Acknowledgments
The idea for this post was based upon the research by Dr. Lloyd Bridge, presented at the British Pharmacological Society meeting December 2018. and published in October 2020 in JPKPD: Hof, F., Bridge, L.J. Exact solutions and equi-dosing regimen regions for multi-dose pharmacokinetics models with transit compartments. J Pharmacokinet Pharmacodyn (2020). https://doi.org/10.1007/s10928-020-09719-8
Introduction
When a PK(/PD) model is developed, it is finally time to put it to use. Many different informative simulations can be performed, one of which is to determine the optimal dosing regimen in which the majority of subjects are within a pre-specified therapeutic window. These simulations can be performed to select the dosing regimen which best balances efficacy with toxicity.
Generally speaking, we can divide these simulations in two approaches: a top-down and bottom-up approach.

With a top-down approach, the clinical applicability determines the ‘degrees of freedom’ you have in your model simulations. For example, a very short dosing interval may never be possible in clinical practice and therefore simulations with intervals of 10 minutes do not have to performed because they will never be used. The same holds true for the ‘optimal’ dose, limitations on the drug formulation can limit the number of possibilities.
At the other side of the spectrum is a bottom-up approach, in which we are interested in the best dose and dosing interval so that the maximal proportion of patients are within a therapeutic window. In order to judge this, we can use a Bayesian approach to estimate the dose and dosing interval but we can also explore a wide range of combinations to judge the overall model behavior.
Therefore, Hof and Bridge et al. introduced the equi-dosing regimen region which gives a guide for selecting therapeutic equi-dosing regimens (JPKPD, Oct 2020). However, even though this article provides a fast analytical solution, it can be expanded with the inter-individual variability that exists in model parameters (differences in clearance, central volume of distribution, etc).
In this post, a script will be given to quickly simulate a range of doses and dosing intervals to judge the optimal dosing within a therapeutic window, using mrgsolve in R, to create the following figure:

Simulation and calculations
The results of a straightforward PK simulation in mrgsolve, or whatever ODE solver is used, is shown below. We have developed our 2-compartment population PK model, included the parameter estimates and simulated a PK profile with inter-individual variability for a certain dose and interval. Click here for more information on this simulation in this post.

If the code to perform such a simulation is new, check out the other posts on pmxsolutions.com.
With the results of this simulation, we can perform a few calculations:
- Subset data from the final interval (assumed to be at steady-state)
- Calculate the minimum and maximum concentration per subject at steady-state
- Determine the % of subjects which minimal and maximal concentrations are:
- Fully within the therapeutic window
- Above the therapeutic minimum
- Below the therapeutic maximum
So how do we do this in R? First we need to define the total simulation time at which our compound has reached steady-state and the dosing interval and dose we would like to test. Let’s say simulate for 120 hours, with a dose of 20 mg every 6 hours.
############################### ## Simulation info sim_time <- 120 ## Time of simulation ii <- 6 ## Interval amt<- 20 ## Amount
Then, we can use our mrgsolve output dataframe (with the individual concentrations per time point) to create a subset and calculate the minimum and maximal concentrations reached per subject at steady-state.
We also want to have a separate dataframe with the simulated dose and dosing interval, which we can easily create from the administration dataframe, to be merged with the results.
## Select the unique combination per individual
combinations <- Administration %>% select(c('ID','amt','ii'))
# Calculate the summary statistics per individual and join with the simulated dose and interval
sum_stat <- df_mrgsolve_output %>%
filter(time >= (sim_time-ii)) %>%
group_by(ID) %>%
summarise(Max_C=max(DV),
Min_C=min(DV)) %>%
left_join(combinations)
Then we need to define the therapeutic window, what are the minimum and maximal concentrations of this compound that we want to obtain?
min_target <- 5 max_target <- 25
We can then calculate the proportion of subjects that fulfill the criteria of being in the target range, above the minimum or below the maximum using the following code.
# sum_stat contains individual results
## Determine if an individual is:
# in range
sum_stat$InTarget <- ifelse(sum_stat$Max_C < max_target & sum_stat$Min_C > min_target,1,0)
# Above minimum
sum_stat$AboveMin <- ifelse(sum_stat$Min_C > min_target,1,0)
# Below maximum
sum_stat$BelowMax <- ifelse(sum_stat$Max_C < max_target,1,0)
grouped <- sum_stat %>%
group_by(amt,ii) %>%
summarise(freq_between=mean(InTarget)*100,
freq_min=mean(AboveMin)*100,
freq_max=mean(BelowMax)*100) %>%
as.data.frame()
This will give us the information that we want! More importantly, we have now reduced the concentration-time profile to only 3 numbers.
However, this is just the result of a single dosing level and dosing interval (20mg every 6 hours). Now lets repeat this 2500x for all different combinations.
Repeating the simulations
First of all, we want to manually set the range of the doses and dosing intervals we would like to explore. We also want to determine the step size (how much doses/intervals), called the resolution in the script below. Furthermore, we need to determine the number of subjects we would like to simulate during each separate simulation. As a general advice, start with a small resolution ~10 and sample size ~100 and check your results.
# Specify the limits of the dose and interval dose_range <-c(10,100) dose_int_range <-c(1,10) resolution = 50 # Number of subjects per simulation n_sim <- 500
We can then create an administration dosing dataframe with all the different combinations we would like to test using the expand.grid() function:
studied_doses <- seq(dose_range[1],dose_range[2],length.out = resolution) studied_intervals <- seq(dose_int_range[1],dose_int_range[2],length.out = resolution) Administration_all <- as.data.frame(expand.grid(cmt=1, time=0, amt=studied_doses, ii=studied_intervals,addl=9999, rate=0,evid=1))
The Administration_all object contains the different combinations we would like to simulate (length = resolution * resolution). Now, we need to initialize our model parameters, and loop through all the different combinations. For this, we can use a simple for loop. But let's use the apply family to do the split-apply-combine strategy instead.
In order to do that, we write the function that applies the calculations to a subset of the Adminstration_all dataframe. It combines all the functions described above. Then we call the apply function on all rows (MARGIN =1) of the Administration_all dataframe and were done!
## Specify the omegas and sigmas matrices
omega <- cmat(etaka,
0, etacl,
0,0,etavd,
0,0,0,etavd2,
0,0,0,0,etaq1)
sigma <- cmat(sigmaprop,
0,sigmaadd)
### Load in model for mrgsolve
mod <- mread_cache("popPK")
## Function to apply to all rows of combinations
calc_summary_pk <- function(dosing_regimen){
## Create a dataframe from a named vector
dosing_regimen <- data.frame(as.list(dosing_regimen))
#### Repeat administration for nsim (multiply the dosing row)
dosing_regimen <- as.data.frame(lapply(dosing_regimen, rep, n_sim))
dosing_regimen <- dosing_regimen %>%
mutate(ID = row_number()) %>% ## Add ID column
arrange(ID,time) ## Sort by ID and time
######################################################################
## Select the unique ID, amt and ii combination of this iteration
combinations <- dosing_regimen %>%
select(c('ID','amt','ii'))
## Set model parameters in dataset, if required. Else mrgsolve model parameters will be used
dosing_regimen$TVKA <- ka
dosing_regimen$TVCL <- cl
dosing_regimen$TVVC <- vd
dosing_regimen$TVVP1 <- vd2
dosing_regimen$TVQ1 <- q1
#################################################################
###### Perform simulation
df_for_analysis <- mod %>%
data_set(dosing_regimen) %>%
omat(omega) %>%
smat(sigma) %>%
mrgsim(start=(sim_time-unique(dosing_regimen$ii)-1),end=sim_time,delta=(unique(dosing_regimen$ii)/50), obsonly=TRUE) %>%
as.data.frame() %>%
left_join(combinations)
## Subset data for the last interval
df_for_analysis <- df_for_analysis %>%
filter(time >= (sim_time-ii))
#################################################################
## Calculate summary statistics
sum_stat <- df_for_analysis %>%
group_by(ID,amt,ii) %>%
summarise(Max_C=max(DV),
Min_C=min(DV))
## Calculate the target concentration binary outcome
# In range
sum_stat$InTarget <- ifelse(sum_stat$Max_C < max_target & sum_stat$Min_C > min_target,1,0)
## Above minimum
sum_stat$AboveMin <- ifelse(sum_stat$Min_C > min_target,1,0)
## Below maximum
sum_stat$BelowMax <- ifelse(sum_stat$Max_C < max_target,1,0)
freq_table <- sum_stat %>%
group_by(amt,ii) %>%
summarise(freq_between=mean(InTarget)*100,
freq_min=mean(AboveMin)*100,
freq_max=mean(BelowMax)*100) %>% as.data.frame()
return(freq_table)
}
############## Execute function over ALL rows (margin=1)
grouped_summary <- apply(Administration_all, MARGIN=1,calc_summary_pk)
## The apply function results are a list. Lets make it a dataframe
grouped_summary <- data.frame(Reduce(rbind,grouped_summary))
We use the apply function instead of performing all simulations in one go due to the exponential increase in the number of simulations when the resolution is increased. This might result in memory issues when millions of profiles are being simulated at once. Trust me, I tried...
Visualization
Our dataframe grouped_summary now contains the information of all different tested combinations of doses and dosing intervals. We can now use the ggplot2 package to add geom_tile() to a figure, in which the color represents the fraction of subjects within the therapeutic window (green = 100%, red = 0%). First, we create a base plot on which we can add the layers that we want to show. Then we can combine the three plots using cowplot's plot_grid() function. We give some extra width to the plot that has the legend.
################################### Graphical output ########################################
## Set base plot to be expanded
base_plot <- ggplot(grouped_summary, aes(amt, ii))+
scale_fill_gradient(low = "red", high = "green",limits = c(0,100))+
scale_x_continuous(name='Dose',limits=c(dose_range[1],dose_range[length(dose_range)]),expand=c(0,0))+
scale_y_continuous(name='Dosing interval',limits=c(dose_int_range[1],dose_int_range[length(dose_int_range)]),expand=c(0,0))+
guides(fill=guide_legend(title='% of subjects'))+
theme(legend.position = "none")+ theme(panel.grid.major = element_blank(), panel.grid.minor = element_blank(), panel.background = element_blank(), axis.line = element_line(colour = "black"))
## Plot with variability within therapeutic minimum
p1 <- base_plot +
geom_tile(aes(fill = freq_between)) +
ggtitle('Subjects within therapeutic window')
## Plot with variability above therapeutic minimum
p2 <- base_plot +
geom_tile(aes(fill = freq_min))+
ggtitle('Subjects above therapeutic minimum')
## Plot with variability below therapeutic maximum
p3 <- base_plot +
geom_tile(aes(fill = freq_min))+
ggtitle('Subjects below therapeutic maximum')+
theme(legend.position = "right") # also include the legend in the final plot
library(cowplot)
plot_grid(p1,p2,p3,ncol=3,rel_widths = c(1,1,1.3))

Percentage of subjects within a therapeutic window, above the therapeutic minimum, and below the therapeutic maximum.
This simulation gives us a perfect overview what the influence is of different dosing regimens on the percentage of subjects within, above, or below a therapeutic window.
This is a quick analysis to see how your model performs in the target therapeutic range and if there are any combinations in which 100% of subjects are within a therapeutic window. For example, in the simulated scenario, a low dose needs to be administered at short intervals to keep everyone between the limits. If dosing intervals of more than 5h are targeted, almost no patients are treated correctly whereas the majority have concentrations below the therapeutic minimum.
The simulation of the 2500x scenarios with a simple PK model takes ~5-10 minutes on a standard Windows desktop. Definitely worth the wait!
The provided code uses the maximum and minimum concentration at steady-state. However, you can implement all sorts of outcomes (AUC, mean, median, Ctrough etc) and create the figures based on that. Additionally, this simulation is not limited to PK, one can also simulate these plots for PD parameters.
Furthermore, if you want to get the information of a single tile in the ggplot, make use of plotly! This will result in an interactive figure that you can hover over. This can be used in the Rstudio Viewer or be incorporated in a shiny app.
library(plotly) ggplotly(p1)

COMMENT
Any suggestions or typo’s? Leave a comment or contact me at info@pmxsolutions.com!
Continue reading:
- From Animals to Algorithms: Leveraging PK/PD Models to Drive the 3R’s in Pharmacology

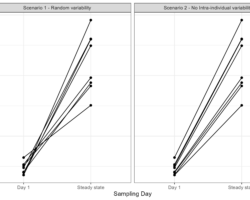
- Exploration of intra-individual variability in a multiple dose study during NCA
- A pharmacometrician’s job interview: the case study

