This blog is an extension on the previously discussed scatter VPC.
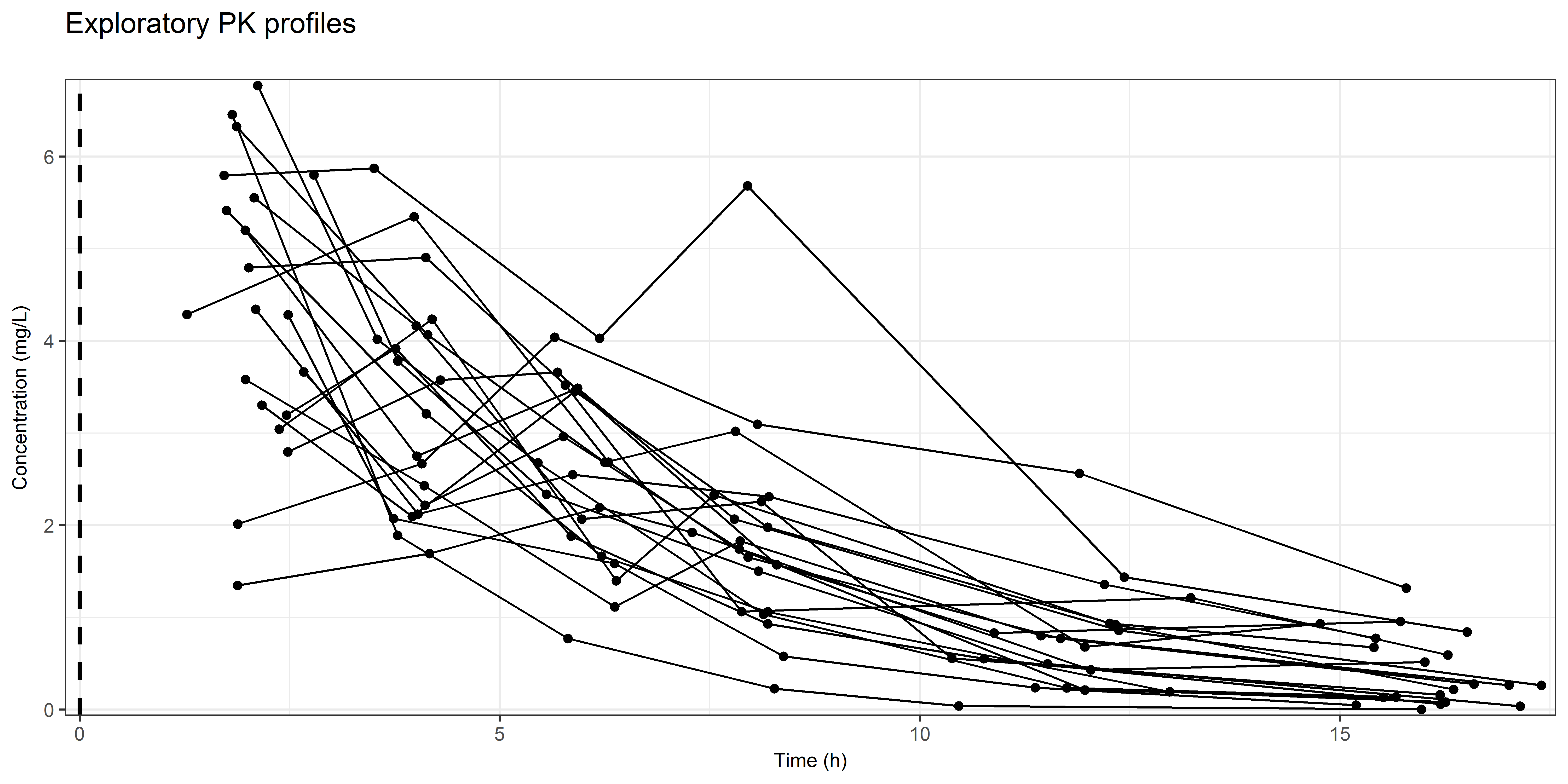
A percentile visual predictive check (VPC) can be used to compare the distribution of the observations with the simulated distributions of your ‘final’ model. In this example, data was generated in which opportunistic sampling around a time point (e.g. 2, 4, 6, 8, 12, 16h) was performed. This results in a scatter of observations around each time point.
Due to this sampling strategy, we need to stratify the observations in bins.
Analysis outline
When we use the PsN function vpc, our dataset used for modelling will be used as the basis for the simulation. This will result in every observation of the dataset that will be simulated 500/1000 times, depending on the chosen number of samples. The no_of_bins and bin_by_count will not be used in this tutorial since we will create our own binning function. In the end, we want to generate the 80% confidence/prediction interval of the observation and the simulated data.
PsN function to create the simulated dataset from Model1:
vpc -samples=500 -no_of_bins=8 -bin_by_count=1 -dir=vpc_Model1 Model1.mod
Getting started
In R, we first want to define the working directory of our ‘final’ model and the prediction interval we want to use at a later stage:
##################################################################################
############################ CODING BLOCK ########################################
##################################################################################
# Run-Time Environment: R version 3.4.0
# Short title: Percentile VPC using ggplot2
#
#
##################################################################################
##################################################################################################
# Clean workspace
rm(list=ls(all=TRUE))## Load some libraries
library(reshape)
library(ggplot2)
library(xpose4)
library(dplyr)#########################################
# Set working directory of model file
setwd("J:\\Model")#########################################
# Set the percentile intervalsPI <- 80 # Specify prediction interval in percentage. Common is 80% interval (10-90%)
# Make a vector for upper and lower percentile to be used in the piping function
perc_PI <- c(0+(1-PI/100)/2, 1-(1-PI/100)/2)
Binning in R
Then, we need to define what kind of bins we want to use (how we specify which observations should be grouped together). We can, for example, divide the bins on different time intervals or by an equal number of observations. In this example we take the easy route, by manually defining the boundaries of the bins around each sampling time point.
#########################################
# Specify the bin times manuallybin_times <- c(0,3,5,7,10,14,18)
Number_of_bins <- length(bin_times)-1
This results in bins between 0-3h, 3-5h, 5-7h, etc. and creates an object which includes the number of bins.
Dataset transformations
When you used the PsN function for the dataset generation, a new folder including a file ending with ‘*1.npctab.dta’ will be created. This is the simulated dataset that we will use, e.g. 500/1000x larger than the original dataset. In the next step we want to load the original dataset and the simulated dataset in our R workspace and do some data cleaning (e.g. removing the dosing rows).
#########################################
######### Read dataset with original observations
Obs <- read.csv("Original_dataset.csv",na.strings = ".")Obs<- Obs[Obs$CMT == 2,] # Remove dosing lines from the dataset
#########################################
## Search for the generated file by PsN in the sub-folders
######### Load in simulated datafiles <- list.files(pattern = "1.npctab.dta", recursive = TRUE, include.dirs = TRUE)
# This reads the VPC npctab.dta file and save it in dataframe_simulations
dataframe_simulations <- read.nm.tables(paste('.\\',files,sep=""))
dataframe_simulations <- dataframe_simulations[dataframe_simulations$MDV == 0,]
After all the datasets are correctly loaded in R, we can start the calculations. We can add a new column BIN to the datasets by reversing a for-loop, dependent on the time. This will start with settings BIN = 6, then 5, 4, etc. It iterates towards the first bin untill all observations have been specified to a BIN.
#########################################
######### Create bins in simulated dataset### Set bins by a backward for-loop
for(i in Number_of_bins:1){
dataframe_simulations$BIN[dataframe_simulations$TIME <= bin_times[i+1]] <-i
}#########################################
######### Create bins in original dataset### Set bins by a backward for loop
for(i in Number_of_bins:1){
Obs$BIN[Obs$TIME <= bin_times[i+1]] <-i
}
We also want to know the median time of each bin, which we can calculate using the following function:
## Get median time of each bin of observations for plotting purposes
bin_middle <- Obs %>%
group_by(BIN) %>%
summarize(bin_middle = median(TIME, na.rm = T))# As a vector
bin_middle <- bin_middle$bin_middle
Calculate the distributions
We can use the stratification column BIN to calculate our summary statistics. This will be done using a piping function for both datasets. Furthermore, the median time points will be added to each row to be eventually used in the plot.
#########################################
######### Calculate percentiles of simulated datasim_vpc <- dataframe_simulations %>%
group_by(BIN) %>%
summarize(C_median = median(DV, na.rm = T),
C_lower = quantile(DV, perc_PI[1], na.rm = T),
C_upper = quantile(DV, perc_PI[2], na.rm = T))sim_vpc$TIME <- bin_middle
#########################################
######### Calculate percentiles of observationsobs_vpc <- Obs %>%
group_by(BIN) %>%
summarize(C_median = median(DV, na.rm = T),
C_lower = quantile(DV, perc_PI[1], na.rm = T),
C_upper = quantile(DV, perc_PI[2], na.rm = T))obs_vpc$TIME <- bin_middle
Graphics
After these calculations, we have all the data that we need! We can specify the lines, points, and theme options that we want in our ggplot2. In this scenario, the red lines are the simulated distributions whereas the black lines are the observations. In the end, we save the image as a .png file with high resolution.
###########################################################################################
########################### Graphics ######################################################
# ----------------------------------------------------------------------------------------#
###########################################################################################Plot_simu <-ggplot()+
###### Simulation percentiles
geom_line(data = sim_vpc,aes(x=TIME,y=C_median),color='red')+
geom_line(data = sim_vpc,aes(x=TIME,y=C_lower),color='red',lty='dashed')+
geom_line(data = sim_vpc,aes(x=TIME,y=C_upper),color='red',lty='dashed')+###### Add observation percentiles
geom_line(data = obs_vpc,aes(x=TIME,y=C_median),color='black')+
geom_line(data = obs_vpc,aes(x=TIME,y=C_lower),color='black',lty='dashed')+
geom_line(data = obs_vpc,aes(x=TIME,y=C_upper),color='black',lty='dashed')+###### Add observations
geom_point(data = Obs,aes(x=TIME,y=DV),color='black')+####### Set ggplot2 theme and settings
theme_bw()+
theme(axis.title=element_text(size=9.0))+
theme(strip.background = element_blank(),
strip.text.x = element_blank(),legend.position="none") +# Set axis labels
labs(x="Time (h)",y="Concentration (mg/L)")+# Add vertical lines to indicate dosing
geom_vline(xintercept = 0, linetype="dashed", size=1) +# Set axis
scale_y_continuous(expand=c(0.01,0))+
scale_x_continuous(expand=c(0.01,0))+## Add title and subtitle
ggtitle("Percentile VPC","")#####################################
## Save as png
png("Figure_Simulation_Percentile_VPC_with_Observations.png",width = 10, height = 5, units='in',res=600)print(Plot_simu)
dev.off()
This analysis will result in the following percentile VPC for our model:

COMMENT
Any suggestions or typo’s? Leave a comment or contact me at info@pmxsolutions.com!